24 April 2026

Nancy J. Cox
1949-2026
It is with profound sadness that we announce the loss of our dear friend and an exceptional leader in global public health, Nancy Cox. As the longtime Head of the U.S. CDC’s Influenza Division and the WHO Collaborating Center for Surveillance, Epidemiology and Control of Influenza, Nancy brought unparalleled dedication, scientific rigor and visionary leadership to the field she loved. Under her guidance, the Global Influenza Surveillance and Response System (GISRS) reached new heights, strengthening the world's ability to detect, monitor, and respond to influenza threats across borders and generations.
Nancy persistent advocacy for transparency and remarkable ability to build consensus among scientists and institutions around the world, proved instrumental in the creation of the GISAID Initiative.
Nancy Cox was truly one of a kind. She will be deeply, dearly missed.
27 February 2026
Recommended composition of influenza virus vaccines for use in the 2026-2027 Northern Hemisphere Influenza Season
(Istanbul, Türkiye) An advisory group of experts taking part in a meeting organized by the WHO Global Influenza Programme between 23-26 February 2026 analyzed influenza virus surveillance data generated by the Global Influenza Surveillance and Response System (GISRS), and issued on 27 February 2026 recommendations on the composition of the influenza vaccines for the next northern hemisphere influenza season.
These recommendations are used by the national vaccine regulatory agencies and the pharmaceutical companies to develop, produce and license influenza vaccines.
15 November 2025
A global collaboration enabled by GISAID data sharing maps H5 influenza antigenic evolution to support vaccine development
Researchers from the University of Cambridge, the University of Wisconsin–Madison, and Erasmus Medical Center are advancing understanding of the rapid genetic and antigenic evolution of H5 influenza viruses. Using genome sequence data shared through GISAID, researchers generated synthetic hemagglutinin constructs, produced recombinant H5 viruses, and performed antigenic characterization to build comprehensive antigenic maps spanning viruses circulating since 1959. These maps guide the design of broadly protective vaccine candidates and support global surveillance.
All antigenic maps are publicly accessible and reagents are available to contributing laboratories, reinforcing the benefits from responsible data sharing, with fair attribution toward equitable global preparedness and response against emerging H5 threats
13 October 2025
Introducing ConvMut, a tool to trace convergent evolution and facilitate variant selection for use in COVID-19 interventions
(Milan/Pisa/Rome) A collaborative development effort by Italian researchers, resulted in a new tool for the analyses of convergent mutations in the coronavirus responsible for COVID-19 to help identify recurrent mutation patterns across different lineages.
The fully integrated ConvMut leverages real-time data in GISAID’s EpiCoV for three kinds of analysis (Convergence Analysis, Codon Exploration, and Mutation Exploration). In the first, central kind of analysis, lineage-specific mutation groups added atop their ancestral nodes are detected, including substitution, deletion and insertion mutations. ConvMut highlights mutations that arise independently in multiple parts of the phylogenetic tree and builds a bar plot to count their representativeness in the tree.
26 September 2025
Recommended composition of influenza virus vaccines for use in the 2026 Southern Hemisphere Influenza Season
(Sapporo, Japan) An advisory group of experts took part in a consultation meeting organized by the WHO Global Influenza Programme between 22-25 September 2025 to analyze influenza virus surveillance data generated by the Global Influenza Surveillance and Response System (GISRS). Their recommendations on the composition of the influenza vaccines for the next Southern hemisphere influenza season were issued on 26 September 2025.
These recommendations are used by the national vaccine regulatory agencies and pharmaceutical companies to develop, produce, and license influenza vaccines. Periodic reevaluation and replacement of viruses on which influenza vaccines are based helps maximize vaccine effectiveness given the dynamic nature of influenza viruses, including those circulating and infecting humans. Recommended reading MMWR Morb Mortal Wkly Rep 2025;74:570–578
21 September 2025
GISAID-GISRS Bioinformatics Workshop in collaboration with the WHO’CC at NIID in the Japan Institute for Health Security
(Sapporo, Japan) On the eve of the vaccine virus selection meeting for the recommended composition of the vaccines for use in the 2026 Southern Hemisphere influenza season, the WHO Collaborating Centre for Reference and Research on Influenza in Tokyo and the Institute for Vaccine Research and Development (IVReD), held a GISAID advanced bioinformatics training workshop on the campus of Hokkaido University, to further harness NGS data for public health responses.
21 April 2025
Data from India's recent fatal human H5N1 case released
On 21 April 2025, the ICMR–National Institute of Virology in Pune, India released specimen data from a recent fatal human H5N1 case in India (EPI_ISL_19836227), belonging to clade 2.3.2.1a and clustering with a 2024 travel-associated case from West Bengal.
Limited surveillance of this lineage in India underscores the urgent need for expanded monitoring in poultry and wild birds to assess its spread, evolution, and zoonotic risk.
28 February 2025
Recommended composition of influenza virus vaccines for use in the 2025-2026 Northern Hemisphere Influenza Season
(London, United Kingdom) An advisory group of experts taking part in a meeting organized by the WHO Global Influenza Programme between 24-27 February 2025 analyzed influenza virus surveillance data generated by the Global Influenza Surveillance and Response System (GISRS), and issued on 28 February 2025 recommendations on the composition of the influenza vaccines for the next northern hemisphere influenza season.
These recommendations are used by the national vaccine regulatory agencies and the pharmaceutical companies to develop, produce and license influenza vaccines.
28 September 2024
GISAID-WHO Training Workshop at Options XII
In collaboration with the WHO's Global Influenza Programme (GIP) and the WHO Collaborating Centre for Reference and Research on Influenza, the GISAID Data Science Initiative held a workshop on the St. Lucia Campus of the University of Queensland on 28 September 2024. The goal is to strengthen the Global Influenza Surveillance and Response System (GISRS) by enhancing genomic surveillance, data sharing, and bioinformatics analysis for respiratory viruses, including influenza, SARS-CoV-2, and RSV.
22 August 2024
Thailand shares first genome of Mpox clade Ib collected on the Asian continent
Thailand's National Institute of Health released the viral genome of the first Asian mpox clade Ib case through GISAID EpiPox (EPI_ISL_19350788), collected from a traveler from the Democratic Republic of the Congo (DRC) on 16-Aug-2024 consistent with its phylogenetic position in the clade I tree. This follows the release of the first European case from Sweden one day earlier.
Since November 2023, the Democratic Republic of the Congo (DRC) has seen a significant increase in mpox cases and the emergence of the new mpox clade Ib variant. Monkeypox virus clade Ib which was first identified in DRC has also been detected in Burundi, Kenya, Rwanda and Uganda.
21 August 2024
Sweden shares first genome of Mpox clade Ib collected outside the African continent
The Public Health Agency of Sweden released through GISAID the viral genome of the first European mpox Clade Ib case that was collected from a traveler from Africa on 13-Aug-2024 (EPI_ISL_19348512).
Since November 2023, the Democratic Republic of the Congo (DRC) has seen a significant increase in mpox virus cases and the emergence of the new mpox Clade Ib variant. The mpox virus Clade Ib, which was first identified in the DRC, has also been detected in neighboring Burundi, Kenya, Rwanda and Uganda. So far, 39 genome sequences of the mpox virus Clade 1b collected in the DRC, Uganda, Kenya and Sweden have been made available on GISAID EpiPox.
24 April 2024
 Brazil and Indonesia conclude the first Arbovirus Summit
Brazil and Indonesia conclude the first Arbovirus Summit
(Bali) Indonesia's Minister of Health, Budi Gunadi Sadikin and Brazil's Minister of Health, Dr. Nísia Trindade Lima convened a two-day summit to gather policy makers, experts in global health and leading researchers of arboviruses, to respond to the significant increase in cases of Dengue virus infections across South America and Southeast Asia. The Arbovirus Summit discussed the urgent need to strengthen integrated arbovirus surveillance and management.
Held at the recently inaugurated GISAID Academy 22-23 April 2024, the Summit showcased innovations in vector control strategies, breakthroughs in antivirals and vaccine development and explore the implementation of global genomic surveillance of arbovirus as a tool for monitoring the effectiveness of intervention strategies as well as viral spread and evolution.
28 September 2023
Since lifting the zero-COVID policy, data from China continues to resemble globally seen patterns
The genomic surveillance of data from China with preliminary phylogenetic analyses over recent months demonstrated a pattern of variant introduction and emergence risk which is substantially similar to that observed globally. Therefore, the daily updates will no longer be shown in the In Focus section. Future GISAID analyses for China will be viewable on the dedicated page accessible here.
Global public health relies on timely genomic surveillance efforts in all countries and regions of the world to detect new evolutionary trends early on.
14 September 2023
16,7 million hCoV-19 sequences shared: Where do surveillance efforts stand?
After mixed efforts in 2020, followed by a massive explosion of Omicron data generated by late 2021 from 215 countries and territories, the number of unique submitting countries, and hence, representativeness of global surveillance, today remains at early 2021 levels, and well above 2020 levels.
While the US, UK, and Germany contributed an outsized share in 2022, it is worth pointing out that increasing representation of diverse regions over time, and the timeliness of submissions are far more critical to ensure new variants are detected in real-time. The recent timely discovery of BA.2.86 and EG.5.1 are examples of continued effectiveness so far.
20 February 2023

H5N1 continues to impact global bird populations
A clade of highly pathogenic avian influenza (HPAI) viruses, originating from poultry but subsequently identified in migratory birds, is severely impacting wild bird populations, including endangered bird species. Over 100 million birds have been killed across the globe, making this the largest outbreak in poultry in history. The current H5N1 clade 2.3.4.4b is also affecting South America where several countries have recorded their first ever cases.
Events with mammal-to-mammal transmission have been reported, including in farmed mink and in marine mammals. Although HPAI H5N1 can cause serious disease in humans, no known human-to-human spread has been identified. GISAID is closely working with its partners to help facilitate the timely sharing of the latest genomic data.
> for more, see statements by WHO Global Influenza Programme and WOAH
10 January 2023
Reflections of a Congolese researcher – three years of COVID-19
Brazzaville, Republic of the Congo - Prof. Dr. Francine Ntoumi looks back on her surveillance work in Africa during the pandemic. In this profile she shares her views on global health equity, lessons learned, the impact of technology, and the importance of international collaboration that flows through GISAID's Regional Hub in Central Africa.
A champion for scientific leadership in Africa and an extraordinary partner, her profile marks the first in a series of features where GISAID is spotlighting the contributions of our diverse community.
10 January 2023
The three-year anniversary of the launch of GISAID’s EpiCoV™
In the early hours of 8 January 2020 at 03:38 PST a short text message sent by the Director General of the China CDC was received by GISAID. These seemingly innocent 25 characters allowed GISAID to prepare before the first whole-genome sequence data would be generated and made available.
On 9 January 2020 at 16:41 PST (10-Jan-2020 00:41 UTC) when these data were shared via GISAID, it kicked off a global surveillance effort of what would become a pandemic of historic proportions. Moreover, it enabled the development of lifesaving countermeasures to COVID-19 at unprecedented speed, including the first vaccines (Polack et al) and the first diagnostic tests (Bohn et al, Carter et al). Three years on, 14.5 million genomes from 215 countries and territories have been shared via GISAID.
6 September 2022
GISAID Training Workshop at Options XI
On the occasion of the 70th Anniversary of GISRS and in collaboration with the World Health Organization, the GISAID Initiative hosted a three-day bioinformatics training workshop at Queen’s University Belfast (27-29 September 2022). The educational programme included instructions on standard and advanced features & functionalities and use of integrated tools for analysis on the GISAID platform.
GISAID also exhibited during Options XI for the Control of Influenza to showcase new developments during the first major in-person gathering of public health professionals since its workshop in Singapore, 2019.
21 June 2022

GISAID joins the G20 Joint Finance & Health Ministerial Meeting
Yogyakarta, 20-21 June. Under the motto ‘Strengthening Global Health Architecture’ a delegation from GISAID attended the G20 Finance and Health Ministers meeting hosted by Indonesia, to discuss with G20 Member States’ and its partners their vision for GISAID+ (GISAID plus).
Global leaders reflected on the need to coordinate and strengthen resources during future pandemics. In particular Member States stressed the essential role GISAID plays in global health security, and discussed how global leaders could support GISAID’s expansion as a global resource to respond to other priority pathogens with major impact and pandemic-potential.
12 April 2022
AHF’s Global Public Health Institute and GISAID team up on genomic sequencing
The AHF Global Public Health Institute at the University of Miami and GISAID announced their collaboration to increase genomic sequencing efforts in low- and middle-Income countries. These efforts are being made possible by a $50,000 donation from AHF to GISAID.
GISAID’s ongoing work in training and skill-sharing will be leveraged to support the genomic sequencing projects AHF is currently funding. GISAID will also work with AHF to advocate for genomic sequencing and sharing as a crucial component of global public health. “Genomic sequencing, in general, is the early warning system that we need if we have additional surges from COVID-19 and other diseases” said AHF President Michael Weinstein.
read the full announcement here
11 April 2022
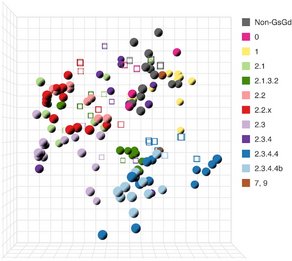
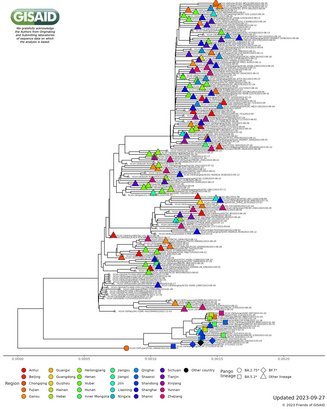
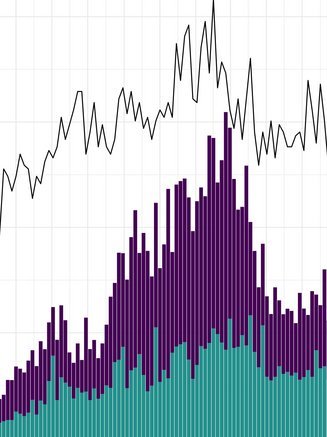
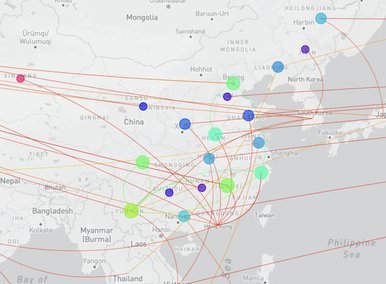
China CDC shares latest COVID-19 data
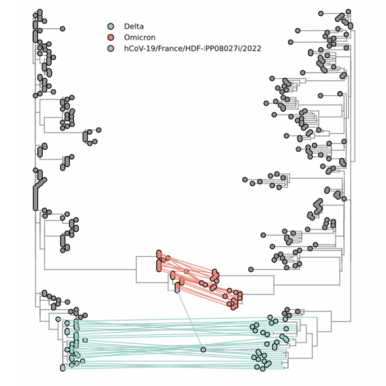
The Chinese Center for Disease Control and Prevention shared via GISAID new hCoV-19 genome sequences from 27 provinces, collected between 2022-01-01 and 2022-03-30. Phylogenetic analyses suggest at least 35 independent introductions from outside China rather than extensive local transmission. This indicates that measures are mostly effective in disrupting long transmission chains following import.
Eleven genomes were assigned as Delta (from six distinct lineages), and 66 genomes were assigned as Omicron (from 14 distinct lineages). No highly divergent new variants were found. A small cluster of multiple cities (2x Hong Kong, 1x Shaanxi, 2x Zhejiang, 2x Yunnan, 1x Beijing) is seen in the tree tip with Hong Kong cases in the genetic base which are part of lineage BA.2.2.
8 March 2022
Confirmation of an AY.4/BA.1 recombinant first detected in France
First solid evidence for a Delta-Omicron recombinant virus has been shared by Institut Pasteur via GISAID (EPI_ISL_10819657 incl. raw reads). The analysis provides definite confirmation of the structure of a recombinant virus derived from the GK/AY.4 and GRA/BA.1 lineages.
This recombinant virus identified in several regions of France by the EMERGEN consortium has been circulating since early January 2022 and genomes with a similar profile have been also identified in Denmark and The Netherlands. Further investigations are needed to determine if these recombinants derive from a single common ancestor or could result from multiple similar recombination events.
25 February 2022 - Recommended composition of influenza virus vaccines for use in the 2022-2023
Recommended composition of influenza virus vaccines for use in the 2022-2023 Northern Hemisphere Influenza Season announced
(Geneva, Switzerland) An advisory group of experts taking part in a virtual meeting organized by the WHO Global Influenza Programme between 21-24 February 2022 analyzed influenza virus surveillance data generated by the WHO Global Influenza Surveillance and Response System (GISRS), and issued on 25 February 2022, recommendations on the composition of the influenza vaccines for the following influenza season.
These recommendations are used by the national vaccine regulatory agencies and the pharmaceutical companies to develop, produce and license influenza vaccines.
24 January 2022
H5 influenza viruses making headlines, again
Europe is experiencing the largest outbreak of highly pathogenic avian influenza A (H5) viruses of clade 2.3.4.4b in poultry for the second year in a row. The outbreaks started with the annual migration of wild birds, including ducks and geese, following the unprecedented year-round virus presence in wild birds during the 2021 summer. The virus was also reported in mammals such as foxes, seals and otters. In December 2021, an outbreak in poultry in Newfoundland, Canada, marked the second arrival of the Eurasian H5 viruses in North America. The virus was subsequently detected in ducks in North and South Carolina, USA.
Distantly related viruses of another subtype A (H5N6), but belonging to the same clade (2.3.4.4b), continue to cause sporadic cases of human infection and fatalities in East and Southeast Asia.
15 January 2022

Mia Brytting
(1964 - 2022)
With deep sadness we announce the loss of our dear friend and member of the scientific community, Mia Brytting. Mia was truly one of a kind and a reliable voice. She made significant contributions and provided scientific guidance during the design of the GISAID platform when it mattered most.
A longstanding member of the GISAID faculty, Mia shared her knowledge of influenza viruses during numerous educational workshops held across the globe since 2009. At the start of COVID-19 pandemic, Mia's guidance and steadfast support helped GISAID rise to the occasion and respond with integrity to fulfill its public health mission. We are going to miss her dearly.
26 November 2021

Omicron discovered on all seven continents
The unique mix of spike amino acid changes in Omicron (clade GRA, lineage B.1.1.529 and descendants BA.1 and BA.2) is of interest as it comprises several that were previously identified to affect receptor binding and antibody escape. As with all low frequency variants with potentially relevant changes, these need to be monitored closely to study if they spread more widely as a consequence of immune escape, or altered receptor interactions. Omicron variants with and without a deletion in spike and a few other changes - BA.1 and BA.2 respectively - are co-circulating, complicating the use of PCR tests to diagnose Omicron based on “S-gene target failure”.
The timely detection of Omicron variants was made possible by researchers from Botswana, Hong Kong, South Africa who shared the first genomes of the variant.
24 September 2021
Recommended composition of influenza virus vaccines for use in the 2022 Southern Hemisphere Influenza Season announced
(Geneva, Switzerland) An advisory group of experts taking part in a virtual meeting organized by the WHO Global Influenza Programme between 13-23 September 2021 analyzed influenza virus surveillance data generated by the WHO Global Influenza Surveillance and Response System (GISRS), and issued on 24 September 2021, recommendations on the composition of the influenza vaccines for the following influenza season.
These recommendations are used by the national vaccine regulatory agencies and the pharmaceutical companies to develop, produce and license influenza vaccines.
22 August 2021
COVID-19 lineages and variants
GISAID’s EpiCoV database employs tools to assign phylogenetic clades and lineages to genetic sequences of the pandemic coronavirus. One such tool is the Pango nomenclature by Rambaut et al (2020) which takes a granular approach to classify and describe viral evolution with detailed lineages.
As new lineages become more widespread, additional genetic markers emerge. Lineage definitions may be updated to allow researchers to track these separately and permit a more fine-grained picture of how a variant is circulating. When these updates occur, all genomes in EpiCoV undergo reclassification by Pango which can lead to temporary fluctuations in the tallies of variants. Overinterpretation of these changes in numbers should be avoided.
10 August 2021

Mamadou Diop
(1986-2021)
The GISAID family mourns the loss of its Senegalese brother and colleague Mamadou Diop. "Mamadou was a well-respected research engineer and bioinformatician at the Institut Pasteur de Dakar", says its Director Dr. Amadou Sall. “His contributions to public health and vision will be missed dearly by all of us.”
Mamadou played an important role in the buildup of GISAID’s first regional hub on the African continent. A native of Dakar, he studied at Lycée Thierno Seydou Nourou Tall and L'Université Gaston Berger de Saint-Louis before joining the Institut Pasteur de Dakar working on software development and bioinformatics pipelines for GWAS and NGS data analyses.
3 June 2021
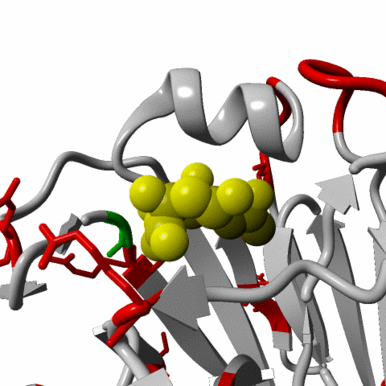
First human case of influenza A/H10N3 virus infection
On April 28, 2021 a resident of the City of Zhenjiang in Jiangsu Province, China, was diagnosed with an influenza A/H10N3 virus infection. This is the first human case of infection with a low pathogenic avian influenza virus of this subtype. How the person was infected is unclear. There is no evidence that the virus spreads easily between humans. A/H10 influenza viruses are abundant in wild birds globally, and occasionally transmit to poultry.
The whole genome sequence of the virus, A/Jiangsu/428/2021, was released rapidly via the GISAID Initiative by the Jiangsu Provincial Center for Disease Control and Prevention in Nanjing (EPI_ISL_2379892).
26 February 2021
Recommended composition of influenza virus vaccines for use in the 2021-2022 Northern Hemisphere Influenza Season announced
(Geneva, Switzerland) An advisory group of experts taking part in a virtual meeting organized by the WHO Global Influenza Programme between 17-25 February 2021 analyzed influenza virus surveillance data generated by the WHO Global Influenza Surveillance and Response System (GISRS), and issued on 26 February 2021, recommendations on the composition of the influenza vaccines for the following influenza season.
These recommendations are used by the national vaccine regulatory agencies and the pharmaceutical companies to develop, produce and license influenza vaccines.
25 February 2021
Human cases of influenza A/H5N8 virus infection
On February 19, 2021 the WHO H5 Reference Laboratory of the State Research Centre for Virology and Biotechnology (VECTOR) of the Russian Federation notified the WHO about cases of human infection with avian influenza A(H5N8) virus. These are the first laboratory confirmed human cases of A(H5N8) virus infection to be reported worldwide. The genome sequences of the viruses from a human case and from poultry are shared in GISAID (EPI_ISL_1038924).
On December 21, 2020, FBRI SRC VB VECTOR Rospotrebnadzor had received 56 serum samples and 37 nasopharyngeal swabs samples from farm workers of Vladimirskaya Poultry Farm in Astrakhan (Russian Federation) who were potentially exposed during a large-scale outbreak of highly pathogenic avian influenza A(H5N8) in poultry.
29 January 2021
Additional closely related bat CoVs identified in Cambodia
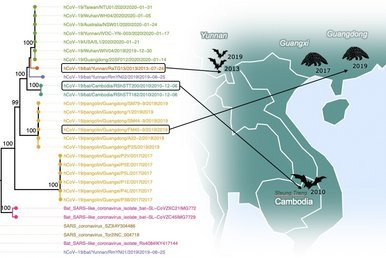
The Virology Unit of the Institut Pasteur du Cambodge has recently submitted the full genome sequences for 2 bat coronaviruses from samples taken from bats in 2010 in the Steung Treng province of Cambodia; they appear to join the list of coronaviruses that are closely related to the current pandemic virus.
Other potential precursors known so far include bat coronaviruses from caves in Yunnan (Southern China), sampled in 2013 and 2019, and samples derived from pangolins in Southern China from 2017 and 2019.
see: Vibol Hul et al (2021) A novel SARS-CoV-2 related coronavirus in bats from Cambodia
10 January 2021
One year since first genomes of SARS-CoV-2 released to the world 10 January 2020 00:41UTC
One year ago, critical public health responses around the globe were kicked off, when China CDC shared via GISAID the first SARS-CoV-2 whole-genomes and associated data.
This curated, high-quality data made available through GISAID permitted the initiation of the development of the first vaccines, diagnostic tests, and other responses at unprecedented speed, including the first vaccines to be approved and made available (Polack et al N Engl J Med 2020), and development of the first NAAT and RT-PCR-based molecular tests to detect the pandemic coronavirus (Bohn et al Clin Chem Lab Med 2020).
22 December 2020
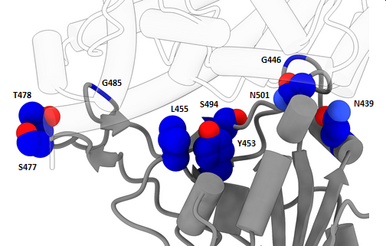
Novel variant 501Y.V2 with triple spike receptor binding site substitutions
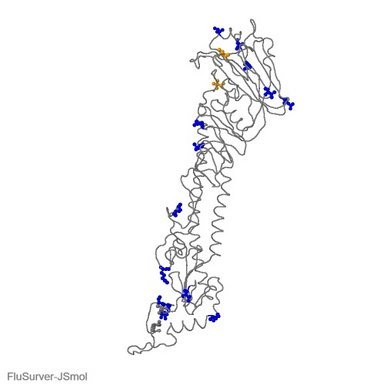
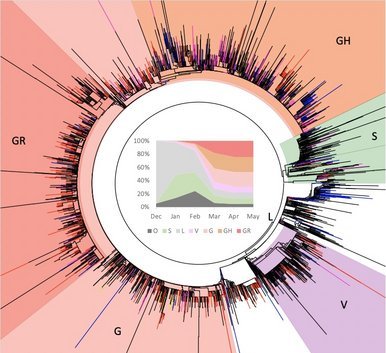
By 21 December, 301 new virus genomes collected in South Africa since November 1 were submitted to GISAID, bringing the total number of virus genome submissions from South Africa to 2,730. The 301 recent virus genomes include 182 from clade GH; 71 from clade GR; 45 from clade G; one from clade GV; and two from other clades. Of the 182 genomes from clade GH, 89% have three mutations in the spike receptor binding site (K417N, E484K, and N501Y). 27% of the 182 have a nine-nucleotide deletion in NSP6 in addition to the three mutations. Spike mutation N501Y is also found in the novel UK variant under investigation, which is from a different clade.
It has been reported, based on high-throughput experiments, that all three spike receptor binding site mutations (K417N, E484K and N501Y) were shown to mildly increase receptor binding. Because receptor binding interfaces are also common epitopes, receptor binding interface mutations could also affect binding of some antibodies to the virus and, in rare cases, have the potential to affect vaccine response. A triple mutant at the interface has not been observed yet in larger outbreaks and should be investigated in detail. Experimental data would be welcome to clarify the impact of the mutations. > read more
21 December 2020
Comment on recent variants and spike protein changes
As seen on many occasions before, mutations are naturally expected for viruses and are most often simply neutral regional markers useful for contact tracing. The changes seen have rarely affected viral fitness and almost never affected clinical outcome but the detailed effects of these mutations remain to be determined fully. Changes in the spike protein have relevance for potential effects on both host receptor as well as antibody binding with possible consequences for infectivity, transmission potential and antibody and vaccine escape. Actual effects need to be measured and verified experimentally.
As has become evident, these few S gene mutations and some deletions are found in multiple genomic contexts (different clades in different countries) that may be an early indication for some potential advantage for these viruses but needs to be verified and does not necessarily mean change in clinical severity or transmission efficiency. > read more
14 December 2020
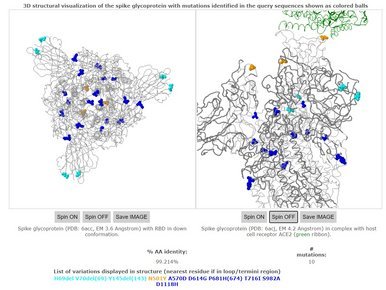
UK reports new variant, termed VUI 202012/01
The United Kingdom reported a new variant, termed VUI 202012/01 (Variant Under Investigation, year 2020, month 12, variant 01). It was defined by multiple spike protein changes (deletion 69-70, deletion 145, N501Y, A570D, D614G, P681H, T716I, S982A, D1118H). An increasing fraction in Southern England (all from clade GR) share several of these changes and a handful have been seen through imports in other countries.
As seen on many occasions before, changes are naturally expected for viruses and are most often simply neutral regional markers useful for contact tracing. The changes seen have rarely been affecting viral fitness and almost never affect clinical outcome but the detailed effects of these changes remain to be determined fully.
30 October 2020
New hCoV-19 variant in Europe, part of natural evolution
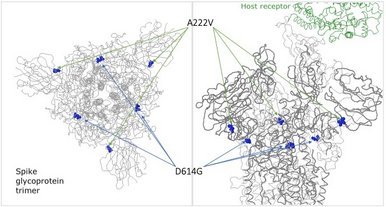
The newest variant of the coronavirus responsible for COVID-19 circulating in Europe features an A222V change in the spike glycoprotein and is designated as clade GV, part of the clade G. The change is not at the receptor binding site but pointing away from it and may, if anything, affect stability of the protein complex with a less pronounced effect than D614G based on its structural position and also not expected to change antibody binding much, depending on the site where the antibody binds.
It's current strong rise within Europe and the age group bias towards young adults could simply point to a founder effect. A222V occurred about 11k times in GISAID submitted genomes (out of 167k) and apart from a handful of exceptions from travel imports, had been restricted to Europe so far.
6 November 2020

Low human health risk of hCoV-19 mutants from mink farms
Mink and other mustelids such as ferrets can be infected relatively easily with hCoV-19. It is not clear why the virus is so contagious in these animals. What is clear is that when viruses infect farmed mink - housed at high densities and in large numbers - this may result in massive virus amplification, posing a high infection risk to humans and animals alike.
During virus amplification in mink, natural selection (“adaptation”) may occur which can be observed by the emergence of mutations that are not seen in human viruses. These mutations are selected because they apparently provide a benefit to the virus in mink. It is important to note that if these mutations would also provide a benefit in humans (resulting in enhanced virulence or transmission) they would have already been naturally selected during infection of (millions of) infected humans hosts. The fact that mink virus variants have not been seen in humans through chains of transmission provides convincing scientific evidence that mutant viruses from mink provide little to none additional health risk to humans.
30 October 2020
First new arrivals of highly pathogenic avian influenza H5 in the 2020-2021 winter in Europe
Highly pathogenic avian influenza viruses of the H5N8 and H5N1 subtypes were detected in the Netherlands in October 2020. The HA genes of the two viruses are closely related, but the other genes of the H5N1 viruses appear to originate from other virus lineages circulating in Eurasia. The earliest virus detections were on 16 October in Eurasian wigeons, a long-distance migrant, and four full virus genomes were uploaded to GISAID by late October.
The virus genome sequences are suggestive for new virus introductions into the EU, different from the outbreaks of last winter. Previously, scientists from the Global Consortium for H5N8 and Related Influenza Viruses used GISAID data to investigate the role of migratory wild birds.
25 September 2020
Recommended composition of influenza virus vaccines for use in the 2021 Southern Hemisphere influenza season announced
Experts taking part in the WHO vaccine composition and information meeting (VCM) between 16-24 September 2020, organized by the WHO Global Influenza Programme, announced on 25. September the recommended components for vaccines for the 2021 Southern Hemisphere influenza season.
These recommendations are used by the national vaccine regulatory agencies and the pharmaceutical companies to develop, produce and license influenza vaccines.
30 August 2020

GISAID wishes Naomi Komadina a Happy 70th Birthday
The Global Initiative congratulates Naomi Komadina, Head of Bioinformatics at WHO Collaborating Centre for Reference and Research on Influenza at the Victorian Infectious Diseases Reference Laboratory (VIDRL) in Melbourne and thanks her for her substantial contributions to GISAID from the start of its incredible journey.
Naomi’s steadfast commitment to responsible data sharing and her technical knowledge passed on to countless participants in GISAID educational workshops around the world, contributed to the global collaboration that enabled GISAID to perform its essential role during outbreaks, such as the new H1N1 influenza virus in 2009 and the emerging coronavirus responsible for the current COVID-19 pandemic.
7 July 2020
New swine flu enters the watchlist
Virus surveillance in animals is an important aspect of pandemic preparedness to know what is out there and could cause zoonotic infections (jumps from animals to humans) or even present pandemic threats. Researchers have concluded a surveillance study in pigs and identified a new reassortant H1N1 swine flu that shows the minimal characteristics for a virus with pandemic potential.
Reassortment of viruses at the animal-human interface continues to be a serious problem. The G4 virus is one of several swine viruses with zoonotic potential in the world that should be watched closely.
4 July 2020
Clade and lineage nomenclature aids in genomic epidemiology studies of active hCoV-19 viruses
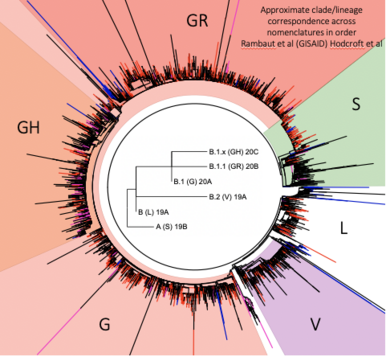
Due to the naturally expanding genetic diversity of hCoV-19 viruses, GISAID introduced a nomenclature system for major clades, developed by Sebastian Maurer-Stroh et al, based on marker mutations within 8 high-level phylogenetic groupings from the early split of S and L, to the further evolution of L into V and G, and later of G into GH, GR and GV, and more recently GR into GRY.
GISAID clades are augmented with more detailed lineages assigned by the Phylogenetic Assignment of Named Global Outbreak LINeages (Pango lineages) tool, aiding in the understanding of patterns and determinants of the global spread of the pandemic strain causing COVID-19. A third effort uses a Year-Letter nomenclature to facilitate discussion of large-scale diversity patterns of hCoV-19 and label clades that persist for at least several months and have significant geographic spread. > read more
13 April 2020
GISAID Comments on the Discussion of Different Types or Clades of the hCoV-19 Virus and their Origin
As the outbreak evolves over time and more data becomes available, several colleagues have analyzed and proposed different "types" of the virus and their origin.
It is important to note that there is currently not enough data from the early outbreak period to interpret the early history of global transmissions from few genomes in detail. Links that seem directly connected now are likely to be connected via other cases also from other countries not sampled and sometimes can be connected differently later with more data.
28 February 2020
Recommended composition of influenza virus vaccines for use in the 2020-2021 Northern Hemisphere Influenza Season announced
(Geneva, Switzerland) An advisory group of experts taking part in a meeting organized by the WHO Global Influenza Programme between 24-27 February 2020 analyzed influenza virus surveillance data generated by the WHO Global Influenza Surveillance and Response System (GISRS), and issued on 28 February 2020, recommendations on the composition of the influenza vaccines for the following influenza season.
These recommendations are used by the national vaccine regulatory agencies and the pharmaceutical companies to develop, produce and license influenza vaccines.
10 Januar 2020
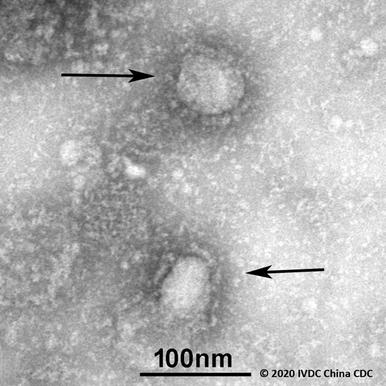
China shares first genome sequences of the newly emerging coronavirus
Laboratories around the world are generating in an unprecedented manner, more and more genome sequences and related clinical and epidemiological data associated with the newly emerging coronavirus (hCoV-19) rapidly made available via GISAID. The pandemic virus was first identified in late December 2019 in Hubei Province, where patients were suffering from respiratory illnesses such as pneumonia. Since then, hCoV-19 is detected across the globe.
The genome sequences of hCoV-19 are crucial to design and evaluate diagnostic tests, to track and trace the ongoing outbreak and to identify potential intervention options.
4 November 2019
Indonesia hosts GISAID Bioinformatics Training Workshop at NIHRD
(Jakarta) Indonesia's National Institute of Health Research and Development (NIHRD) and GISAID held a influenza bioinformatics training workshop from 4-8 November. Hands-on training was provided by GISAID DTG members from the WHO Collaborating Centre for Reference and Research on Influenza, Melbourne and the Bioinformatics Institute (BII), Singapore.
Participants from across Indonesia were trained on a new computational pipeline introduced by BII to generate consensus sequences from next generation sequencing data, and on the GISAID platform to monitor complex epidemiological relationships among circulating animal and human influenza viruses and the detection and interpretation of evolutionary changes. see the Report
1 November 2019
World Flu Day 2019: China’s Campaign for Flu Awareness and Academic Conference
The Chinese Center for Disease Control and Prevention hosted in Beijing the launch of the Popular Science Promotion Campaign for Flu Awareness & Academic Conference in Celebration of the 2nd World Flu Day on 1. November 2019.
Under the motto 'Know Flu, Prevent Flu, Beat Flu' researchers agreed to unite in a push for stronger political will across the globe, calling for the continuing support of influenza prevention and control. The event welcomed 800 participants who reflected on the progress made since the 2009 H1N1 Pandemic.
> see Xinhua News (2019) and The Lancet (2018)
30 September 2019

Vicki Gregory
(1968-2019)
With deep sadness we announce the loss of Vicki Gregory, a member of our scientific family and dear friend. Vicki was a founding member of GISAID’s Database Technical Group playing a key role in providing scientific guidance for the development of GISAID’s EpiFlu database.
Vicki was held in high regard by colleagues around the world for her contribution to influenza surveillance and research. A wonderful mother, wife, daughter and friend to many, Vicki inspired us all.
27 September 2019
Recommended composition of influenza virus vaccines for use in the 2020 Southern Hemisphere influenza season announced
(Geneva) Experts taking part in the WHO vaccine composition and information meeting (VCM) between 23-25 September 2019, organized by the WHO Global Influenza Programme, announced today the recommended components for vaccines for the 2020 Southern Hemisphere influenza season.
These recommendations are used by the national vaccine regulatory agencies and the pharmaceutical companies to develop, produce and license influenza vaccines.
27 August 2019
GISRS - GISAID Bioinformatics Training Workshop with leading experts in Singapore
The WHO Global Influenza Programme, together with the National Centre for Infectious Diseases and the Bioinformatics Institute Singapore and the GISAID Initiative held a 2-day influenza bioinformatics workshop with hands-on sessions provided by leading experts in their respective fields, preceding the Options for the Control of Influenza X on 26-27 August 2019.
Participants trained on the GISAID EpiFlu™ database to monitor complex epidemiological relationships among circulating animal and human influenza viruses and the detection and interpretation of evolutionary change using FluSurver and Nextflu.
23 May 2019
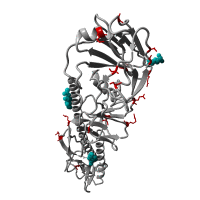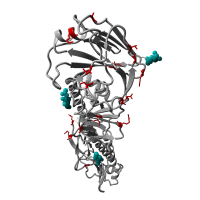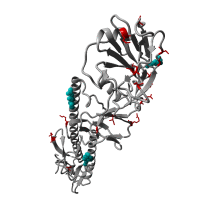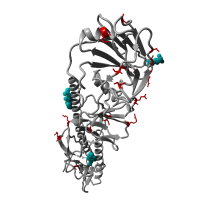
Re-emergence of highly pathogenic avian influenza H7N9 viruses in humans in mainland China, in 2019
A severe human case of highly pathogenic avian influenza (HPAI) H7N9 occurred in late March 2019 in the Inner Mongolia autonomous region of China. The case resided near a poultry slaughtering stall, prompting immediate environmental sampling of this location.
Phylogenetic analyses of the virus showed that the sequences from the case and from several environmental samples formed a distinct clade from the gene sequences from viruses collected in 2017. Comparison with a candidate vaccine strain showed several mutations, possibly associated with antigenic drift which has yet to be characterized. No human cases of H7N9 were reported for over a year previously.
13 May 2019
GISAID Comments on the WHO Report on the Public Health Implications of Implementation of the Nagoya Protocol
Member States representatives met at the 72nd annual World Health Assembly in Geneva, Switzerland to discuss and debate a report prepared by the WHO on the public health implications of implementation the Nagoya Protocol.
GISAID comments on that paper, highlighting the current issues around sharing of seasonal influenza viruses, the consequences of delays in virus sharing, and the connection to the discussion at the Convention on Biological Diversity.
23 April 2019

Leading Flu Expert at US' CDC Dr. Katz says goodbye
(Atlanta) After nearly three decades at the US CDC, the Director of the WHO Collaborating Center for Surveillance, Epidemiology and Control of Influenza and Co-Chair of GISAID Scientific Advisory Council, Dr. Jackie Katz has retired. Since joining the CDC in 1992, Dr. Katz drove progress in influenza-related science and public health. She is the recipient of three Charles C. Shepard Science Awards, having co-authored over 330 peer-reviewed research articles reviews, and book chapters.
“While Dr. Katz’s leadership was most evident during the 2009 flu pandemic, her most lasting legacy may be her ability to develop young scientists through mentorship and support.” said Dr. Dan Jernigan, Director of the CDC's Influenza Division
29 March 2019

Prominent Influenza Researcher and Director of Japan's NIID bids farewell
(Tokyo) Dr. Takato Odagiri, Director of the Influenza Virus Research Center at the National Institute of Infectious Diseases (NIID) and Director of the WHO Collaborating Center for Reference and Research on Influenza has retired.
Dr. Odagiri began his career in 1985 as an assistant professor and a lecturer at Jichi Medical University and is the author and co-author of over 100 peer-reviewed publications.
Dr. Odagiri served on GISAID's Scientific Advisory Council since 2014, playing a critical role in providing scientific oversight of the Initiative. GISAID wishes Dr. Odagiri a healthy & happy new chapter in his life and a successful retirement.
1 November 2018

World Flu Day: momentum from China for influenza control
The 1st World Flu Day was formally launched at the Asian-Pacific Centenary Spanish 1918-flu symposium in Shenzhen, to raise public awareness of influenza and accelerate scientific innovation and basic research efforts particularly the development of a universal flu vaccine. Prof George Fu Gao, Director of the Chinese Center for Disease Control and Prevention and fellow researchers are pushing for stronger global political will in continuing the support of influenza prevention and control.
“2018 also marks the 15-year commemoration of the SARS outbreak, after which strengthening the CDC became the top priority in China's public policy agenda. As a result, China boosted investment in the public health system, strengthening national and local surveillance systems for all infectious diseases more efficiently and effectively, and improving research capacity, especially for emerging infectious diseases.” says Prof. Gao
18 October 2018
Marie-Paule Kieny talks Ethics and Data Sharing at Grand Challenges 2018
Dr. Marie-Paule Kieny spoke in Berlin on October 18th at the Grand Challenges Annual Meeting, co-hosted by the Bill & Melinda Gates Foundation and the German Federal Ministry of Education & Research, discussing "Sequence data sharing – the influenza GISAID model and its wider applicability".
“In a world where global emerging disease threats demand effective sharing of information, GISAID has demonstrated what can be achieved by galvanizing the interests of public health officials, academia and industry to combat influenza” says Dr Kieny
8 October 2018
GIHSN to share epidemiological and virological data on severe influenza via GISAID
Sanofi Pasteur’s Foundation for Influenza Epidemiology has awarded GISAID €300,000 to strengthen the Initiative and enhance its EpiFlu™ database and educational programs. In partnership with the Foundation’s Global Influenza Hospital-based Surveillance Network (GIHSN), GISAID will help promote the sharing and integration of clinical, epidemiological and virological data to better understand the impact of severe influenza and the benefit of vaccination.
“Given the high variability of influenza virus circulation, generation of comparable data across seasons with a broad geographical scope are required”, says Dr. Cédric Mahé, President of the Foundation and Head of Epidemiology at Sanofi Pasteur.
28 September 2018

Dr. Karin Schwabenbauer, Germany's Chief Veterinary Officer, a former President of the OIE World Assembly of Delegates and OIE Council, and FAO official overseeing the Animal Health Crisis Management Centre retires at the end of September. As the German Government's focal point in the GISAID Initiative, she played a key role in ensuring GISAID's recognition in FAO, OIE and WHO member states deliberations following the 2009 H1N1 pandemic.
“Karin's passion for science and unwavering commitment to improving both animal and human health made her one of the true champions of One Health. We thank Karin for her unswerving support of our Initiative, she has helped shape.” said Dr. Alan Hay, GISAID Scientific Liaison Officer
27 September 2018
Recommended composition of influenza virus vaccines for use in the 2019 Southern Hemisphere Influenza Season announced
(Atlanta) Experts taking part in the WHO vaccine composition and information meeting (VCM) between 24-26 September, at the WHO Collaborating Centre for Surveillance, Epidemiology & Control of Influenza at the US Centers for Disease Control and Prevention (CDC), recommend components for vaccines for the 2019 Southern Hemisphere influenza season.
> read more:
26 July 2018

GISAID announces a major contribution from Seqirus
GISAID welcomes a €250,000 donation from Seqirus, a leading innovator in influenza vaccine technologies and pandemic response solutions. Seqirus is the first influenza vaccine company to make a significant financial contribution to GISAID.
“Seqirus is proud to support GISAID in its mission to provide open and rapid access to influenza virus data for the public health good of all nations,” said Gordon Naylor, President of Seqirus
> read more:
21 May 2018

Renowned scientist and public health specialist to advise the GISAID Initiative on international affairs
Dr. Marie-Paule Kieny, a former Assistant Director-General of WHO will advise GISAID on international affairs. Most recently Dr Kieny led the WHO research & development initiative during the Ebola crisis resulting in the WHO R&D Blueprint for public health emergencies, with an emphasis on enhanced global collaboration and timely sample and data sharing to support rapid emergency response.
“In a world where global emerging disease threats demand effective sharing of information, GISAID has demonstrated what can be achieved by galvanizing the interests of public health officials, academia and industry to combat influenza” said Dr. Kieny
22 March 2018

GISAID-WHO Bioinformatics Training Workshop
Madagascar, 22 - 23 March 2018
During the 6th African Network for the Surveillance of Influenza (ANISE) in Antananarivo, GISAID and isirv - in collaboration with Institut Pasteur de Madagascar and the World Health Organization’s Global Influenza Programme (GIP), with support from United States Agency for International Development (USAID) - are holding a two-day training workshop on genetic sequence analysis of influenza viruses to provide training for monitoring sequence variation among influenza viruses in relation to surveillance of influenza epidemics and detection of resistance to influenza antiviral drugs.
17 July 2017

Celebrating 70 Years GIP and 65 Years Global Influenza Surveillance and Response System
GISAID salutes GIP and GISRS. It was 1947 when the WHO’s Interim Committee recognized the importance of Influenza and started a globally-coordinated effort for its surveillance, study and control that saw the beginning of the Global Influenza Programme. By 1952, the WHO Executive Board called for an influenza surveillance system to collect, correlate and distribute information regarding occurrence, epidemiology and laboratory findings. Today GISRS encompasses 143 institutions across 113 Member States. A global disease surveillance network built on voluntary collaboration and real-time reporting that makes up the backbone of today's global influenza surveillance.
19 May 2017
G20 Health Ministers recognize the importance of GISAID in regard to virus data sharing
At the invitation of Germany, the first meeting of Health Ministers of the Group of Twenty leading industrialized and emerging economies (G20) took place in Berlin between 19-20 May 2017.
Under the banner of “Together Today for a Healthy Tomorrow – Joint Commitment for Shaping Global Health”, the two-day meeting focused on combating global health hazards. In their Berlin Declaration, the G20 Health Ministers recognize the importance of the Global Initiative on Sharing All Influenza Data (GISAID).
> read more
> The Berlin Declaration of the G20 Health Ministers (2017)
1 August 2017

Editorial Board affirms Open Access designation of GISAID
re3data.org and DataCite, the world’s leading provider of digital object identifiers (DOI) for research data, affirmed the designation of access to GISAID's database and data as Open Access. A persistent link for this designation has been assigned to GISAID for citation purposes (doi:10.17616/R3Q59F).
GISAID satisfies all conditions of the definition for open access contributions of the Berlin Declaration on Open Access to Knowledge in the Sciences and Humanities, last but not least, due to GISAID's ability "to provide a mechanism to enforce proper attribution“ through its user identification procedures, validating terms of use and licenses.
> http://doi.org/10.17616/R3Q59F
> The Berlin Declaration on Open Access to Knowledge in the Sciences and Humanities (2003)
2 March 2017
GISAID SAC Informal Consultation on Collaboration with GISRS
WHO Headquarters, 2 March 2017
Directors of WHO Collaborating Centers and Members of GISAID's Scientific Advisory Council consulted with the Global Influenza Program, National Influenza Centers and ERLs, as well as industry representatives on the progress of timely virus data sharing through GISAID, followed by round-table discussions.
26 February 2015
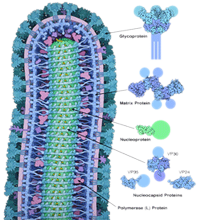
Sharing of Ebola data needs good practice framework, what GISAID did for influenza
The rapid dissemination of results during outbreaks is sporadic at best. In the case of influenza, a global initiative called GISAID established a framework for good practice. Largely thanks to this, during the 2009 H1N1 influenza outbreak, it became a go-to place for the community to deposit and locate H1N1 sequence information.
Pardis C. Sabeti et al, Nature 518, 477–479 (26 February 2015)
11 October 2011
WHO is fully supportive of GISAID
During the 1st GISAID Symposium hosted by the Federal Republic of Germany, Assistant Director General Dr. Keiji Fukuda expresses WHO's support for GISAID.
"We have the development of critically important and technically advanced new platforms such as GISAID. This data sharing initiative provides an important option for sharing genetic sequence and epidemiological data. WHO is fully supportive of GISAID and any other initiative which promotes sharing and access to information, in ways that are trustworthy, transparent, efficient and timely."